Single-cell omics (SCO) has revolutionized the way and resolution by which life science research is conducted. SCO encompasses multiple technologies that are able to profile various omic modalities at the single-cell level. The rapid development of these technologies has been accompanied by the active development of data analysis methods.
The goal of the Single-cell omics data analysis club events is to bring together people in the field to discuss leading-edge data analysis approaches. Each event has one topic that will be introduced by a guest speaker, followed by a discussion. The club is open to everyone, and the participation of method developers and single-cell bioinformatics heavy users is particularly encouraged. The events are organized by ELIXIR-FI (CSC) and the Biocenter Finland Bioinformatics Platform.
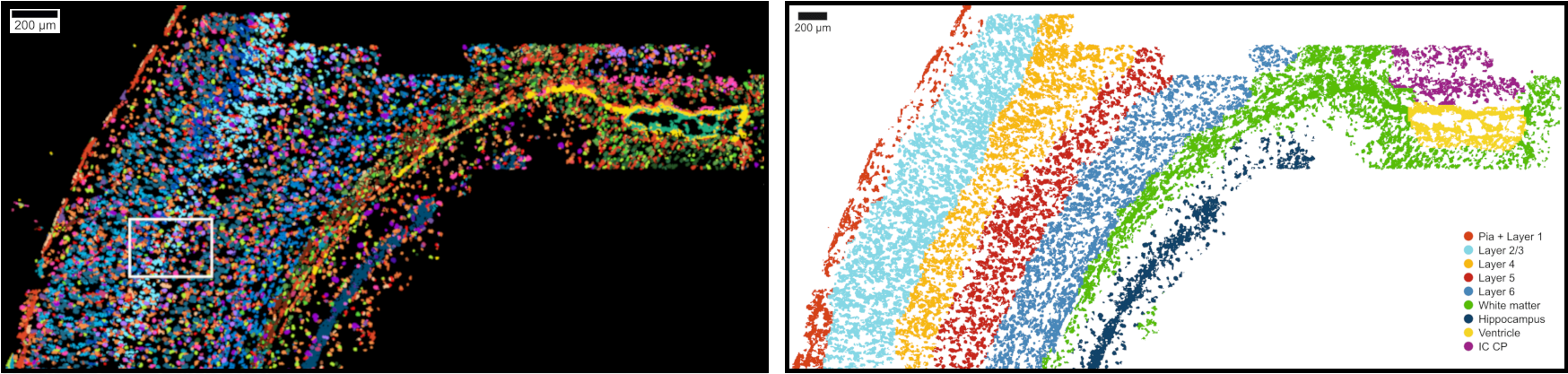
The topic of this first club event is spatial transcriptomics data analysis and it is introduced by Naveed Ishaque (Charite Berlin).
Naveed is group leader at BIH@Charité - Center for Digital Health. His Computational Oncology group works on spatially resolved transcriptomics, and they have recently published a tool called SSAM - the first computational framework for segmentation free cell-type and tissue domain analysis of in situ transcriptomics data. References:
- Park et al (2021): Cell segmentation-free inference of cell types from in situ transcriptomics data. Nature communications, 12(1), 3545.
- Tiesmeyer et al (2022): SSAM-lite: A Light-Weight Web App for Rapid Analysis of Spatially Resolved Transcriptomics Data. Front Genet. 2022 Feb 28;13:785877.